WGAN-GP with R-GCN 用于生成小分子图
作者: akensert
创建日期 2021/06/30
最后修改日期 2021/06/30
描述: WGAN-GP with R-GCN 的完整实现,用于生成新颖的分子。
简介
在本教程中,我们将实现一个用于图的生成模型,并使用它来生成新颖的分子。
动机:新药(分子)的开发可能非常耗时且成本高昂。深度学习模型的应用可以通过预测已知分子的性质(例如,溶解度、毒性、对目标蛋白的亲和力等)来缓解对良好候选药物的搜索。由于可能分子的数量是天文数字,我们搜索/探索分子的空间仅占整个空间的一小部分。因此,实现可以学习生成新颖分子(否则将永远不会被探索到)的生成模型是可取的。
参考文献(实现)
本教程中的实现基于/受到 MolGAN 论文 和 DeepChem 的 Basic MolGAN 的启发。
延伸阅读(生成模型)
近期用于分子图的生成模型的实现还包括 Mol-CycleGAN、GraphVAE 和 JT-VAE。有关生成对抗网络的更多信息,请参阅 GAN、WGAN 和 WGAN-GP。
设置
安装 RDKit
RDKit 是一个用 C++ 和 Python 编写的化学信息学和机器学习软件集合。在本教程中,RDKit 用于方便高效地将 SMILES 转换为分子对象,然后从这些对象中获取原子和键的集合。
SMILES 以 ASCII 字符串的形式表示给定分子的结构。SMILES 字符串是一种紧凑的编码,对于较小的分子来说,它相对人类可读。将分子编码为字符串既可以缓解又可以促进对给定分子的数据库和/或网络搜索。RDKit 使用算法准确地将给定的 SMILES 转换为分子对象,然后可以使用该对象计算大量的分子性质/特征。
请注意,RDKit 通常通过 Conda 安装。但是,由于 rdkit_platform_wheels 的存在,RDKit 现在(在本教程中)可以通过 pip 轻松安装,如下所示:
pip -q install rdkit-pypi
为了方便可视化分子对象,还需要安装 Pillow:
pip -q install Pillow
导入包
from rdkit import Chem, RDLogger
from rdkit.Chem.Draw import IPythonConsole, MolsToGridImage
import numpy as np
import tensorflow as tf
from tensorflow import keras
RDLogger.DisableLog("rdApp.*")
数据集
本教程使用的数据集是来自 MoleculeNet 的 量子力学数据集 (QM9)。尽管数据集中附带了许多特征和标签列,但我们只关注 SMILES 列。QM9 数据集是生成图的良好入门数据集,因为分子中发现的最大重原子(非氢原子)数量仅为九个。
csv_path = tf.keras.utils.get_file(
"qm9.csv", "https://deepchemdata.s3-us-west-1.amazonaws.com/datasets/qm9.csv"
)
data = []
with open(csv_path, "r") as f:
for line in f.readlines()[1:]:
data.append(line.split(",")[1])
# Let's look at a molecule of the dataset
smiles = data[1000]
print("SMILES:", smiles)
molecule = Chem.MolFromSmiles(smiles)
print("Num heavy atoms:", molecule.GetNumHeavyAtoms())
molecule
SMILES: Cn1cncc1O
Num heavy atoms: 7
定义辅助函数
这些辅助函数将有助于将 SMILES 转换为图,并将图转换为分子对象。
分子图的表示。分子可以自然地表示为无向图 G = (V, E),其中 V 是顶点(原子)集,E 是边(键)集。对于此实现,每个图(分子)将表示为一个邻接张量 A,该张量编码原子对的存在/不存在,并具有一个用于表示键类型的 one-hot 编码扩展维度;以及一个特征张量 H,对于每个原子,它使用 one-hot 编码表示其原子类型。请注意,由于 RDKit 可以推断出氢原子,因此为了简化模型,将氢原子从 A 和 H 中排除。
atom_mapping = {
"C": 0,
0: "C",
"N": 1,
1: "N",
"O": 2,
2: "O",
"F": 3,
3: "F",
}
bond_mapping = {
"SINGLE": 0,
0: Chem.BondType.SINGLE,
"DOUBLE": 1,
1: Chem.BondType.DOUBLE,
"TRIPLE": 2,
2: Chem.BondType.TRIPLE,
"AROMATIC": 3,
3: Chem.BondType.AROMATIC,
}
NUM_ATOMS = 9 # Maximum number of atoms
ATOM_DIM = 4 + 1 # Number of atom types
BOND_DIM = 4 + 1 # Number of bond types
LATENT_DIM = 64 # Size of the latent space
def smiles_to_graph(smiles):
# Converts SMILES to molecule object
molecule = Chem.MolFromSmiles(smiles)
# Initialize adjacency and feature tensor
adjacency = np.zeros((BOND_DIM, NUM_ATOMS, NUM_ATOMS), "float32")
features = np.zeros((NUM_ATOMS, ATOM_DIM), "float32")
# loop over each atom in molecule
for atom in molecule.GetAtoms():
i = atom.GetIdx()
atom_type = atom_mapping[atom.GetSymbol()]
features[i] = np.eye(ATOM_DIM)[atom_type]
# loop over one-hop neighbors
for neighbor in atom.GetNeighbors():
j = neighbor.GetIdx()
bond = molecule.GetBondBetweenAtoms(i, j)
bond_type_idx = bond_mapping[bond.GetBondType().name]
adjacency[bond_type_idx, [i, j], [j, i]] = 1
# Where no bond, add 1 to last channel (indicating "non-bond")
# Notice: channels-first
adjacency[-1, np.sum(adjacency, axis=0) == 0] = 1
# Where no atom, add 1 to last column (indicating "non-atom")
features[np.where(np.sum(features, axis=1) == 0)[0], -1] = 1
return adjacency, features
def graph_to_molecule(graph):
# Unpack graph
adjacency, features = graph
# RWMol is a molecule object intended to be edited
molecule = Chem.RWMol()
# Remove "no atoms" & atoms with no bonds
keep_idx = np.where(
(np.argmax(features, axis=1) != ATOM_DIM - 1)
& (np.sum(adjacency[:-1], axis=(0, 1)) != 0)
)[0]
features = features[keep_idx]
adjacency = adjacency[:, keep_idx, :][:, :, keep_idx]
# Add atoms to molecule
for atom_type_idx in np.argmax(features, axis=1):
atom = Chem.Atom(atom_mapping[atom_type_idx])
_ = molecule.AddAtom(atom)
# Add bonds between atoms in molecule; based on the upper triangles
# of the [symmetric] adjacency tensor
(bonds_ij, atoms_i, atoms_j) = np.where(np.triu(adjacency) == 1)
for (bond_ij, atom_i, atom_j) in zip(bonds_ij, atoms_i, atoms_j):
if atom_i == atom_j or bond_ij == BOND_DIM - 1:
continue
bond_type = bond_mapping[bond_ij]
molecule.AddBond(int(atom_i), int(atom_j), bond_type)
# Sanitize the molecule; for more information on sanitization, see
# https://www.rdkit.org/docs/RDKit_Book.html#molecular-sanitization
flag = Chem.SanitizeMol(molecule, catchErrors=True)
# Let's be strict. If sanitization fails, return None
if flag != Chem.SanitizeFlags.SANITIZE_NONE:
return None
return molecule
# Test helper functions
graph_to_molecule(smiles_to_graph(smiles))
生成训练集
为了节省训练时间,我们将仅使用 QM9 数据集的十分之一。
adjacency_tensor, feature_tensor = [], []
for smiles in data[::10]:
adjacency, features = smiles_to_graph(smiles)
adjacency_tensor.append(adjacency)
feature_tensor.append(features)
adjacency_tensor = np.array(adjacency_tensor)
feature_tensor = np.array(feature_tensor)
print("adjacency_tensor.shape =", adjacency_tensor.shape)
print("feature_tensor.shape =", feature_tensor.shape)
adjacency_tensor.shape = (13389, 5, 9, 9)
feature_tensor.shape = (13389, 9, 5)
模型
其思想是实现一个通过 WGAN-GP 的生成器网络和一个判别器网络,最终得到一个可以生成小的新颖分子(小图)的生成器网络。
生成器网络需要能够将(批次中的每个示例)向量 z 映射到 3 维邻接张量(A)和 2 维特征张量(H)。为此,z 将首先通过一个全连接网络,然后其输出将通过两个独立的全连接网络。这两个全连接网络中的每个网络将(为批次中的每个示例)输出一个 tanh 激活向量,然后进行重塑和 softmax 操作,以匹配多维邻接/特征张量。
由于判别器网络将接收来自生成器或训练集的图(A、H)作为输入,因此我们需要实现图卷积层,这使得我们能够操作图。这意味着输入到判别器网络的将首先通过图卷积层,然后是平均池化层,最后是几个全连接层。最终输出应该是每批次一个标量,指示相关输入的“真实性”(在这种情况下是“假”或“真”分子)。
图生成器
def GraphGenerator(
dense_units, dropout_rate, latent_dim, adjacency_shape, feature_shape,
):
z = keras.layers.Input(shape=(LATENT_DIM,))
# Propagate through one or more densely connected layers
x = z
for units in dense_units:
x = keras.layers.Dense(units, activation="tanh")(x)
x = keras.layers.Dropout(dropout_rate)(x)
# Map outputs of previous layer (x) to [continuous] adjacency tensors (x_adjacency)
x_adjacency = keras.layers.Dense(tf.math.reduce_prod(adjacency_shape))(x)
x_adjacency = keras.layers.Reshape(adjacency_shape)(x_adjacency)
# Symmetrify tensors in the last two dimensions
x_adjacency = (x_adjacency + tf.transpose(x_adjacency, (0, 1, 3, 2))) / 2
x_adjacency = keras.layers.Softmax(axis=1)(x_adjacency)
# Map outputs of previous layer (x) to [continuous] feature tensors (x_features)
x_features = keras.layers.Dense(tf.math.reduce_prod(feature_shape))(x)
x_features = keras.layers.Reshape(feature_shape)(x_features)
x_features = keras.layers.Softmax(axis=2)(x_features)
return keras.Model(inputs=z, outputs=[x_adjacency, x_features], name="Generator")
generator = GraphGenerator(
dense_units=[128, 256, 512],
dropout_rate=0.2,
latent_dim=LATENT_DIM,
adjacency_shape=(BOND_DIM, NUM_ATOMS, NUM_ATOMS),
feature_shape=(NUM_ATOMS, ATOM_DIM),
)
generator.summary()
Model: "Generator"
__________________________________________________________________________________________________
Layer (type) Output Shape Param # Connected to
==================================================================================================
input_1 (InputLayer) [(None, 64)] 0
__________________________________________________________________________________________________
dense (Dense) (None, 128) 8320 input_1[0][0]
__________________________________________________________________________________________________
dropout (Dropout) (None, 128) 0 dense[0][0]
__________________________________________________________________________________________________
dense_1 (Dense) (None, 256) 33024 dropout[0][0]
__________________________________________________________________________________________________
dropout_1 (Dropout) (None, 256) 0 dense_1[0][0]
__________________________________________________________________________________________________
dense_2 (Dense) (None, 512) 131584 dropout_1[0][0]
__________________________________________________________________________________________________
dropout_2 (Dropout) (None, 512) 0 dense_2[0][0]
__________________________________________________________________________________________________
dense_3 (Dense) (None, 405) 207765 dropout_2[0][0]
__________________________________________________________________________________________________
reshape (Reshape) (None, 5, 9, 9) 0 dense_3[0][0]
__________________________________________________________________________________________________
tf.compat.v1.transpose (TFOpLam (None, 5, 9, 9) 0 reshape[0][0]
__________________________________________________________________________________________________
tf.__operators__.add (TFOpLambd (None, 5, 9, 9) 0 reshape[0][0]
tf.compat.v1.transpose[0][0]
__________________________________________________________________________________________________
dense_4 (Dense) (None, 45) 23085 dropout_2[0][0]
__________________________________________________________________________________________________
tf.math.truediv (TFOpLambda) (None, 5, 9, 9) 0 tf.__operators__.add[0][0]
__________________________________________________________________________________________________
reshape_1 (Reshape) (None, 9, 5) 0 dense_4[0][0]
__________________________________________________________________________________________________
softmax (Softmax) (None, 5, 9, 9) 0 tf.math.truediv[0][0]
__________________________________________________________________________________________________
softmax_1 (Softmax) (None, 9, 5) 0 reshape_1[0][0]
==================================================================================================
Total params: 403,778
Trainable params: 403,778
Non-trainable params: 0
__________________________________________________________________________________________________
图判别器
图卷积层。 关系图卷积层 实现非线性变换的邻域聚合。我们可以这样定义这些层:
H^{l+1} = σ(D^{-1} @ A @ H^{l+1} @ W^{l})
其中 σ 表示非线性变换(通常是 ReLU 激活),A 是邻接张量,H^{l} 是第 l 层的特征张量,D^{-1} 是 A 的逆对角度数张量,W^{l} 是第 l 层的可训练权重张量。具体来说,对于每种键类型(关系),度数张量在其对角线上表示连接到每个原子的键的数量。请注意,在本教程中,D^{-1} 被省略了,原因有两个:(1)在这种归一化如何应用于生成器生成的连续邻接张量尚不清楚;(2)没有归一化的 WGAN 的性能似乎效果不错。此外,与 原始论文 不同,没有定义自环,因为我们不希望训练生成器预测“自键合”。
class RelationalGraphConvLayer(keras.layers.Layer):
def __init__(
self,
units=128,
activation="relu",
use_bias=False,
kernel_initializer="glorot_uniform",
bias_initializer="zeros",
kernel_regularizer=None,
bias_regularizer=None,
**kwargs
):
super().__init__(**kwargs)
self.units = units
self.activation = keras.activations.get(activation)
self.use_bias = use_bias
self.kernel_initializer = keras.initializers.get(kernel_initializer)
self.bias_initializer = keras.initializers.get(bias_initializer)
self.kernel_regularizer = keras.regularizers.get(kernel_regularizer)
self.bias_regularizer = keras.regularizers.get(bias_regularizer)
def build(self, input_shape):
bond_dim = input_shape[0][1]
atom_dim = input_shape[1][2]
self.kernel = self.add_weight(
shape=(bond_dim, atom_dim, self.units),
initializer=self.kernel_initializer,
regularizer=self.kernel_regularizer,
trainable=True,
name="W",
dtype=tf.float32,
)
if self.use_bias:
self.bias = self.add_weight(
shape=(bond_dim, 1, self.units),
initializer=self.bias_initializer,
regularizer=self.bias_regularizer,
trainable=True,
name="b",
dtype=tf.float32,
)
self.built = True
def call(self, inputs, training=False):
adjacency, features = inputs
# Aggregate information from neighbors
x = tf.matmul(adjacency, features[:, None, :, :])
# Apply linear transformation
x = tf.matmul(x, self.kernel)
if self.use_bias:
x += self.bias
# Reduce bond types dim
x_reduced = tf.reduce_sum(x, axis=1)
# Apply non-linear transformation
return self.activation(x_reduced)
def GraphDiscriminator(
gconv_units, dense_units, dropout_rate, adjacency_shape, feature_shape
):
adjacency = keras.layers.Input(shape=adjacency_shape)
features = keras.layers.Input(shape=feature_shape)
# Propagate through one or more graph convolutional layers
features_transformed = features
for units in gconv_units:
features_transformed = RelationalGraphConvLayer(units)(
[adjacency, features_transformed]
)
# Reduce 2-D representation of molecule to 1-D
x = keras.layers.GlobalAveragePooling1D()(features_transformed)
# Propagate through one or more densely connected layers
for units in dense_units:
x = keras.layers.Dense(units, activation="relu")(x)
x = keras.layers.Dropout(dropout_rate)(x)
# For each molecule, output a single scalar value expressing the
# "realness" of the inputted molecule
x_out = keras.layers.Dense(1, dtype="float32")(x)
return keras.Model(inputs=[adjacency, features], outputs=x_out)
discriminator = GraphDiscriminator(
gconv_units=[128, 128, 128, 128],
dense_units=[512, 512],
dropout_rate=0.2,
adjacency_shape=(BOND_DIM, NUM_ATOMS, NUM_ATOMS),
feature_shape=(NUM_ATOMS, ATOM_DIM),
)
discriminator.summary()
Model: "model"
__________________________________________________________________________________________________
Layer (type) Output Shape Param # Connected to
==================================================================================================
input_2 (InputLayer) [(None, 5, 9, 9)] 0
__________________________________________________________________________________________________
input_3 (InputLayer) [(None, 9, 5)] 0
__________________________________________________________________________________________________
relational_graph_conv_layer (Re (None, 9, 128) 3200 input_2[0][0]
input_3[0][0]
__________________________________________________________________________________________________
relational_graph_conv_layer_1 ( (None, 9, 128) 81920 input_2[0][0]
relational_graph_conv_layer[0][0]
__________________________________________________________________________________________________
relational_graph_conv_layer_2 ( (None, 9, 128) 81920 input_2[0][0]
relational_graph_conv_layer_1[0][
__________________________________________________________________________________________________
relational_graph_conv_layer_3 ( (None, 9, 128) 81920 input_2[0][0]
relational_graph_conv_layer_2[0][
__________________________________________________________________________________________________
global_average_pooling1d (Globa (None, 128) 0 relational_graph_conv_layer_3[0][
__________________________________________________________________________________________________
dense_5 (Dense) (None, 512) 66048 global_average_pooling1d[0][0]
__________________________________________________________________________________________________
dropout_3 (Dropout) (None, 512) 0 dense_5[0][0]
__________________________________________________________________________________________________
dense_6 (Dense) (None, 512) 262656 dropout_3[0][0]
__________________________________________________________________________________________________
dropout_4 (Dropout) (None, 512) 0 dense_6[0][0]
__________________________________________________________________________________________________
dense_7 (Dense) (None, 1) 513 dropout_4[0][0]
==================================================================================================
Total params: 578,177
Trainable params: 578,177
Non-trainable params: 0
__________________________________________________________________________________________________
WGAN-GP
class GraphWGAN(keras.Model):
def __init__(
self,
generator,
discriminator,
discriminator_steps=1,
generator_steps=1,
gp_weight=10,
**kwargs
):
super().__init__(**kwargs)
self.generator = generator
self.discriminator = discriminator
self.discriminator_steps = discriminator_steps
self.generator_steps = generator_steps
self.gp_weight = gp_weight
self.latent_dim = self.generator.input_shape[-1]
def compile(self, optimizer_generator, optimizer_discriminator, **kwargs):
super().compile(**kwargs)
self.optimizer_generator = optimizer_generator
self.optimizer_discriminator = optimizer_discriminator
self.metric_generator = keras.metrics.Mean(name="loss_gen")
self.metric_discriminator = keras.metrics.Mean(name="loss_dis")
def train_step(self, inputs):
if isinstance(inputs[0], tuple):
inputs = inputs[0]
graph_real = inputs
self.batch_size = tf.shape(inputs[0])[0]
# Train the discriminator for one or more steps
for _ in range(self.discriminator_steps):
z = tf.random.normal((self.batch_size, self.latent_dim))
with tf.GradientTape() as tape:
graph_generated = self.generator(z, training=True)
loss = self._loss_discriminator(graph_real, graph_generated)
grads = tape.gradient(loss, self.discriminator.trainable_weights)
self.optimizer_discriminator.apply_gradients(
zip(grads, self.discriminator.trainable_weights)
)
self.metric_discriminator.update_state(loss)
# Train the generator for one or more steps
for _ in range(self.generator_steps):
z = tf.random.normal((self.batch_size, self.latent_dim))
with tf.GradientTape() as tape:
graph_generated = self.generator(z, training=True)
loss = self._loss_generator(graph_generated)
grads = tape.gradient(loss, self.generator.trainable_weights)
self.optimizer_generator.apply_gradients(
zip(grads, self.generator.trainable_weights)
)
self.metric_generator.update_state(loss)
return {m.name: m.result() for m in self.metrics}
def _loss_discriminator(self, graph_real, graph_generated):
logits_real = self.discriminator(graph_real, training=True)
logits_generated = self.discriminator(graph_generated, training=True)
loss = tf.reduce_mean(logits_generated) - tf.reduce_mean(logits_real)
loss_gp = self._gradient_penalty(graph_real, graph_generated)
return loss + loss_gp * self.gp_weight
def _loss_generator(self, graph_generated):
logits_generated = self.discriminator(graph_generated, training=True)
return -tf.reduce_mean(logits_generated)
def _gradient_penalty(self, graph_real, graph_generated):
# Unpack graphs
adjacency_real, features_real = graph_real
adjacency_generated, features_generated = graph_generated
# Generate interpolated graphs (adjacency_interp and features_interp)
alpha = tf.random.uniform([self.batch_size])
alpha = tf.reshape(alpha, (self.batch_size, 1, 1, 1))
adjacency_interp = (adjacency_real * alpha) + (1 - alpha) * adjacency_generated
alpha = tf.reshape(alpha, (self.batch_size, 1, 1))
features_interp = (features_real * alpha) + (1 - alpha) * features_generated
# Compute the logits of interpolated graphs
with tf.GradientTape() as tape:
tape.watch(adjacency_interp)
tape.watch(features_interp)
logits = self.discriminator(
[adjacency_interp, features_interp], training=True
)
# Compute the gradients with respect to the interpolated graphs
grads = tape.gradient(logits, [adjacency_interp, features_interp])
# Compute the gradient penalty
grads_adjacency_penalty = (1 - tf.norm(grads[0], axis=1)) ** 2
grads_features_penalty = (1 - tf.norm(grads[1], axis=2)) ** 2
return tf.reduce_mean(
tf.reduce_mean(grads_adjacency_penalty, axis=(-2, -1))
+ tf.reduce_mean(grads_features_penalty, axis=(-1))
)
训练模型
为了节省时间(如果在 CPU 上运行),我们将仅训练模型 10 个 epoch。
wgan = GraphWGAN(generator, discriminator, discriminator_steps=1)
wgan.compile(
optimizer_generator=keras.optimizers.Adam(5e-4),
optimizer_discriminator=keras.optimizers.Adam(5e-4),
)
wgan.fit([adjacency_tensor, feature_tensor], epochs=10, batch_size=16)
Epoch 1/10
837/837 [==============================] - 197s 226ms/step - loss_gen: 2.4626 - loss_dis: -4.3158
Epoch 2/10
837/837 [==============================] - 188s 225ms/step - loss_gen: 1.2832 - loss_dis: -1.3941
Epoch 3/10
837/837 [==============================] - 199s 237ms/step - loss_gen: 0.6742 - loss_dis: -1.2663
Epoch 4/10
837/837 [==============================] - 187s 224ms/step - loss_gen: 0.5090 - loss_dis: -1.6628
Epoch 5/10
837/837 [==============================] - 187s 223ms/step - loss_gen: 0.3686 - loss_dis: -1.4759
Epoch 6/10
837/837 [==============================] - 199s 237ms/step - loss_gen: 0.6925 - loss_dis: -1.5122
Epoch 7/10
837/837 [==============================] - 194s 232ms/step - loss_gen: 0.3966 - loss_dis: -1.5041
Epoch 8/10
837/837 [==============================] - 195s 233ms/step - loss_gen: 0.3595 - loss_dis: -1.6277
Epoch 9/10
837/837 [==============================] - 194s 232ms/step - loss_gen: 0.5862 - loss_dis: -1.7277
Epoch 10/10
837/837 [==============================] - 185s 221ms/step - loss_gen: -0.1642 - loss_dis: -1.5273
<keras.callbacks.History at 0x7ff8daed3a90>
使用生成器采样新颖分子
def sample(generator, batch_size):
z = tf.random.normal((batch_size, LATENT_DIM))
graph = generator.predict(z)
# obtain one-hot encoded adjacency tensor
adjacency = tf.argmax(graph[0], axis=1)
adjacency = tf.one_hot(adjacency, depth=BOND_DIM, axis=1)
# Remove potential self-loops from adjacency
adjacency = tf.linalg.set_diag(adjacency, tf.zeros(tf.shape(adjacency)[:-1]))
# obtain one-hot encoded feature tensor
features = tf.argmax(graph[1], axis=2)
features = tf.one_hot(features, depth=ATOM_DIM, axis=2)
return [
graph_to_molecule([adjacency[i].numpy(), features[i].numpy()])
for i in range(batch_size)
]
molecules = sample(wgan.generator, batch_size=48)
MolsToGridImage(
[m for m in molecules if m is not None][:25], molsPerRow=5, subImgSize=(150, 150)
)
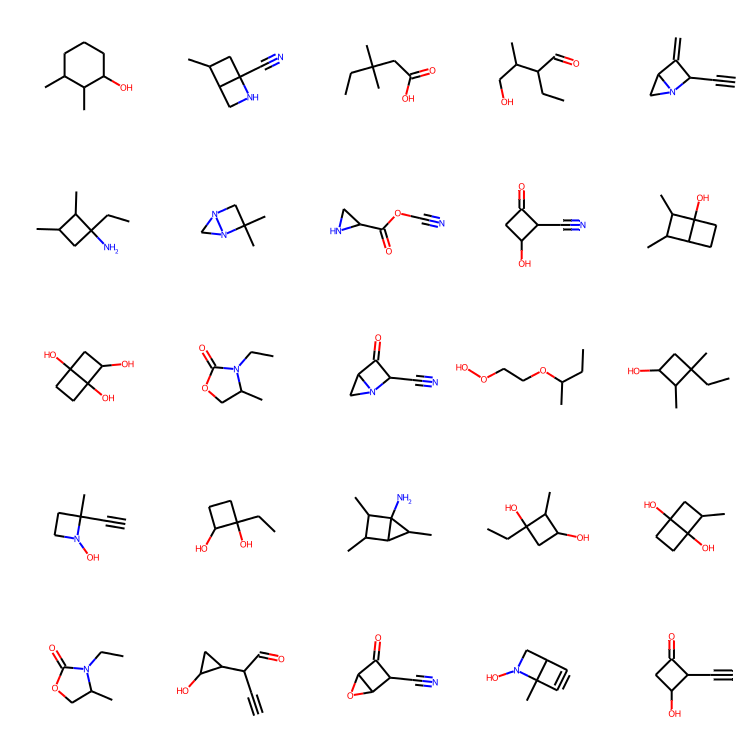
结论
检查结果。 训练十个 epoch 似乎足以生成一些看起来不错的分子!请注意,与 MolGAN 论文 相比,本教程中生成分子的独特性似乎非常高,这很好!
我们学到的东西和前景。在本教程中,我们成功地实现了一个用于分子图的生成模型,它使我们能够生成新颖的分子。未来,实现可以修改现有分子的生成模型(例如,优化现有分子的溶解度或蛋白质结合性)将很有趣。为此,可能需要一个重构损失,但由于没有简单明了的方法来计算两个分子图之间的相似性,因此实现起来很棘手。
HuggingFace 上提供的示例
| 训练好的模型 | 演示 |
|---|---|
 |
 |